Blog, genetics, potato (Solanum tuberosum)
Potato: Color Genetics
A Summary of Potato Color Genetics
| Introduction | |
| Genes Controlling Color | |
| Pigments | |
| Purple and Red | |
| Yellow and Orange | |
| Color Distribution | |
| Flower Color | |
| Foliage Color | |
| Russeting | |
| Known Genotypes | |
Introduction
For many crops, you don’t have to search very far to find a nice summary chart of the major genes involved in economically important traits. This kind of information makes breeding decisions a lot easier. Knowledge of the genetics underlying traits allows the breeder to identify varieties that are homozygous for a particular trait and to use those varieties to produce predictable progeny. Unfortunately, no such thing exists for potatoes as far as I have seen. This surprised me, because the potato is one of the most studied of all food crops. However, potato breeding also doesn’t have the same kind of following among amateurs as do crops like tomatoes, peppers, and corn, so there is less demand for easily digestible information.

That leaves the scientific literature and there is plenty of it. Unfortunately, as is the case with the taxonomy of potatoes, there are multiple systems that have changed with time and that also have been applied differently for diploids and tetraploids. I’ve done my best to pull together the most current information here in a way that unifies the treatments of both diploids and tetraploids. I’m taking the approach suggested by De Jong (1991), of using the diploid classification scheme (Dodds 1955, 1956) in preference to the earlier tetraploid scheme (Salaman 1910) wherever possible. Where there are popular alternatives, I’ve noted them. Diploids and tetraploids have the same genes, but some of them may not function in the same way in tetraploids, where additional copies of the gene can result in the trait almost always or almost never being expressed, or being expressed in an intermediate form. For a review of how ploidy can affect gene expression in potatoes, see Potato: Genetics – The Basics.
Potatoes make life complicated for the breeder in several ways. Many potato traits are quantitative rather than qualitative. Qualitative traits are well represented by classical, Mendelian loci. Quantitative traits are determined by the interactions of complexes of quantitative trait loci (QTLs). What’s the difference? At the biochemical level, these are really the same thing, but the outcome that the breeder experiences is quite different. A Mendellian locus controls a single trait in a dominant/recessive regime. A good example of this in potato is the P gene. Dominant P- turns on purple anthocyanidin production and recessive pp turns it off. While you may not get purple tubers even with dominant P due to the involvement of distribution genes, you will never get purple tubers with recessive pp. A QTL is usually part of a group of loci that collectively determine a particular trait. It might be necessary to have several different genes all in the right state in order for color to be expressed at all, or the genes might be additive, each affecting the intensity of the color. In some cases, the qualitative genes described here are supplemented by other quantitative genes, which usually are not as well understood and certainly not as easy for the breeder to work with.

Another problem is the presence of multiallelic loci. Multiallelic loci contain three or more alleles and therefore cannot be analyzed in the binary fashion used for the more common biallelic loci. B is an example of a multiallelic locus, with five alleles identified to date. Typically, these alleles operate in a winner take all fashion, with the highest valued allele taking the dominant position, but unusual things can happen with increased dosage in polyploids.
It is likely that I have made mistakes and, particularly, oversimplifications in this post. The relationships between diploid and tetraploid genetics are not always as clear as I have made them out to be. The evidence for some genes is better than others. Informed by my experience, which is not exhaustive, where I had to make a choice between improving the practical usability of this post or emphasizing uncertainty, I have generally chosen the former. I have proceeded under the assumption that diploid and tetraploid genetics must be the same and forced the systems to line up accordingly. For a more cautious review of potato color genetics, see De Jong (1991).
Genes Controlling Color
There are two main types of color genes: those that control the production of a pigment and those that control the distribution of a pigment. The following table summarizes the main qualitative color genes in the potato and their interactions.
Summary of Qualitative Loci Operating in Diploid and Tetraploid Potatoes
| Locus | Alleles | Chr. | Function | Notes | References |
| Ac | Ac/ac | Acylation of anthocyanins. When present in homozygous recessive configuration (acac), the color of the flowers is a deeper red or blue color than otherwise. | Harborne 1960 | ||
| B | Bd/Bc/Bb/Ba/b | 10 | Controls distribution of pigmentation to flower abscission zone (Ba-d), eyebrow (Bb-d), seed spot (Bc,d), and petiole base (Bd). Recessive bb blocks color distribution to all of these locations. |
Only functions with dominant P or R, as pigments are otherwise not produced, so cannot be distributed. Bc and Bd have been observed only in diploids (Dodds 1962). |
Dodds 1956 |
| I (D) | I/isp/i (D/d) | 10 | Controls distribution of color to tuber skin. Intensity of color may be in part dosage dependent on I, so tetraploids can display wider gradation. The genotype Iisp produces spectacled tubers with non-pigmented regions around the eyes (Dodds 1962). | I is often used when discussing diploids and D for tetraploids. Because tetraploids are more common, D is more prevalent in the literature. E/e has also been used and appears to be nearly synonymous. In diploids, I is epistatic to Pf; neither skin nor flesh color will develop with recessive ii. Sometimes divided into Ico and Iep, for cortex and epidermis, respectively, although this doesn’t appear to be commonly used. |
I: Dodds 1956 D: Salaman 1910 D: Howard 1970 |
| F | F/f | 10 | Distribution of color to flower. Homozygous recessive configuration (ff) causes reduced distribution of color to the flowers. | Only functions with dominant P or R, as pigments are otherwise not produced, so cannot be distributed. | |
| M | M/m | ? | Restricts skin pigmentation to the eyes when I is dominant. | Some sources consider this dubious, although it does explain colored eyes in some tetraploids that have white skin but dominant P or R. | Howard 1970 |
| P | P/p | 11 | Controls purple/blue pigmentation through the productions of anthocyanins (petunidin derivatives). P- produces color, pp does not. | P is epistatic to R; more purple pigments are produced than red, even when R is also dominant. It is important to note that red pigments are not turned off, just produced in lesser amounts. | |
| Pd | Pd/pd | ? | Pigmented dorsal side. Controls distribution of red and purple pigements to the top sides of leaves. | Used with diploids; unclear how it applies to tetraploids. | Garg 1981 |
| Pf | Pf/pf | ? | Pigmented flesh. Controls distribution of red and purple pigments to tuber flesh. | Requires dominant I (D). There are at least a few varieties that have colored flesh but not skin, which clearly does not fit this system. Perhaps there is another gene involved that suppresses distribution to the skin in those cases. I suspect that this gene is also responsible for anther color. | De Jong 1987 |
| Pv | Pv/pv (Ul/ul) | 10 | Pigmented ventral side. Controls distribution of red and purple pigments to undersides of leaves | Pv is used with diploids and Ul with tetraploids, but they appear to be the same thing. |
Pv: Garg 1981 Ul: Kessel 1974 |
| Pw | Pw/pw | 10 | Pigmented whorl. The young foliage, particularly at emergence, shows red or purple pigmentation. |
Used with diploids; unclear how it applies to tetraploids. The effect of this gene is best observed in newly emerging foliage. |
Kessel 1974 |
| R | R/Rpw/r | 2 | Controls red pigmentation through the production of anthocyanins (pelargonidin derivatives). R- produces color, while rr does not. When the Rpw allele is present in homozygous configuration, the plant produces peonanin instead of peonidin, and this results in pink color rather than red (Howard 1970). |
Also previously described as D. Rpw suppresses cyanidin and produces white flowers (Dodds 1962), but I presume only when homozygous. |
|
| Vc |
Vc/vc |
4 | Controls pigmentation of leaf veins, petioles, and stems. |
This locus was discovered in the wild potato Solanum venturii. It is unknown whether it functions in domesticated potatoes, but it seems likely. Presumably, color is only produced by this locus when P is also dominant. |
|
| Y | Y/y | 3 | Controls the production of yellow flesh and, presumably, skin as well. | Controls production of the pigment lutein. The allele O (orange) was previously included at this locus, but more recent evidence has shown that to be incorrect. | |
| Ym | Ym/ym | Yellow margin. When homozygous recessive (ymym), plants have a yellow margin around the leaves and a glossy appearance. | There are also structural changes: leaves are smaller than usual and some plants are dwarfed. | Dodds 1962 | |
| Zep | Zep*/Zep1 | Controls the production of orange flesh and, presumably, skin as well. | Controls production of the pigment zeaxanthin in combination with Chy2. | Wolters 2010 |
Genes that are located on the same chromosome may be inherited together. The major color distribution genes are all on chromosome 10 and are often inherited as a group. This is known as linkage. It isn’t certain that they will be inherited together because recombination swaps large portions of a chromosome in the process of sexual reproduction, but linkage is always possible when genes are located on the same chromosome.
The following table shows example genotypes for different tuber skin and flesh color combinations. I almost added this table when I first wrote this article, but decided against it because it is fraught with peril. Some of these genotypes are probably right in theory but wrong in practice. It is also almost certain that I have made mistakes in here. But, many of the questions that I have received about the article center on how the different genes fit together to produce different phenotypes, so I think it is worth the risk of total humiliation and rejection by my peers.
The dash in some genotypes is a wildcard. So, for example, in a diploid, if you see P-, that means PP or Pp. In a tetraploid P- could be PPPP, PPPp, PPpp, or Pppp. Wildcards tell you that a dominant allele is at work.
| Phenotype | Diploid Genotype(s) | Tetraploid Genotype(s) | |
| White tuber skin |
ii pp rr yy I- |
iiii pppp rrrr yyyy I— |
|
| With white flesh |
ii pp rr yy I- |
iiii pppp rrrr yyyy I— |
|
| With red spectacles | pp R- I- M- | pppp R— I— M— | |
| With blue spectacles | P- I- M- | P— I— M— | |
| With red eyebrows | pp R- ii Bb-d– | pppp R— iiii Bb-d— | |
| With blue eyebrows | P- ii Bb-d– | P— iiii Bb-d— | |
| Blue tuber skin | P- I- | P— I— | |
| With white spectacles | P- Iisp | P— Iisp— | |
| With yellow spectacles | P- Y- Iisp | P— Y— Iisp— | |
| With blue flesh | P- I- Pf- | P— I— Pf— | |
| With white flesh | P- I- pfpf | P— I— pfpfpfpf | |
| With yellow flesh | P- I- Y- pfpf | P— I— Y— pfpfpfpf | |
| Red tuber skin | pp R- I- | pppp R— I— | |
| With white spectacles | pp R- Iisp | pppp R— Iisp— | |
| With yellow spectacles | pp R- Y- Iisp | pppp R— Y— Iisp— | |
| With red flesh | pp R- I- Pf- | pppp R— I— Pf— | |
| With white flesh | pp R- I- pfpf | pppp R— I— pfpfpfpf | |
| With yellow flesh | pp R- I- Y- pfpf | pppp R— I— Y— pfpfpfpf | |
| Yellow tuber skin |
ii Y- pp rr Y- |
iiii Y— pppp rrrr Y— |
|
| With yellow flesh |
pp rrI- Y- ii Y- |
pppp rrrr I— Y— iiii Y— |
|
| With orange flesh |
pp rr I- Chy23– Zep1Zep1 ii Chy23– Zep1Zep1 |
pppp rrrr I— Chy23— Zep1Zep1Zep1Zep1 iiii Chy23— Zep1Zep1Zep1Zep1 |
|
| With red spectacles | pp R- I- M- Y- | pppp R— I— M— Y— | |
| With blue spectacles | P- I- M- Y- | P— I— M— Y— | |
| With red eyebrows | pp R- ii Bb-d– Y- | pppp R— iiii Bb-d— Y— | |
| With blue eyebrows | P- ii Bb-d– Y- | P— iiii Bb-d— Y— | |
| Pink tuber skin | pp Rpw– I- | pppp Rpw— I— | |
| With white flesh | pp Rpw– I- | pppp Rpw— I— | |
| With yellow flesh | pp Rpw– I- Y- | pppp Rpw— I— Y— | |
Pigments
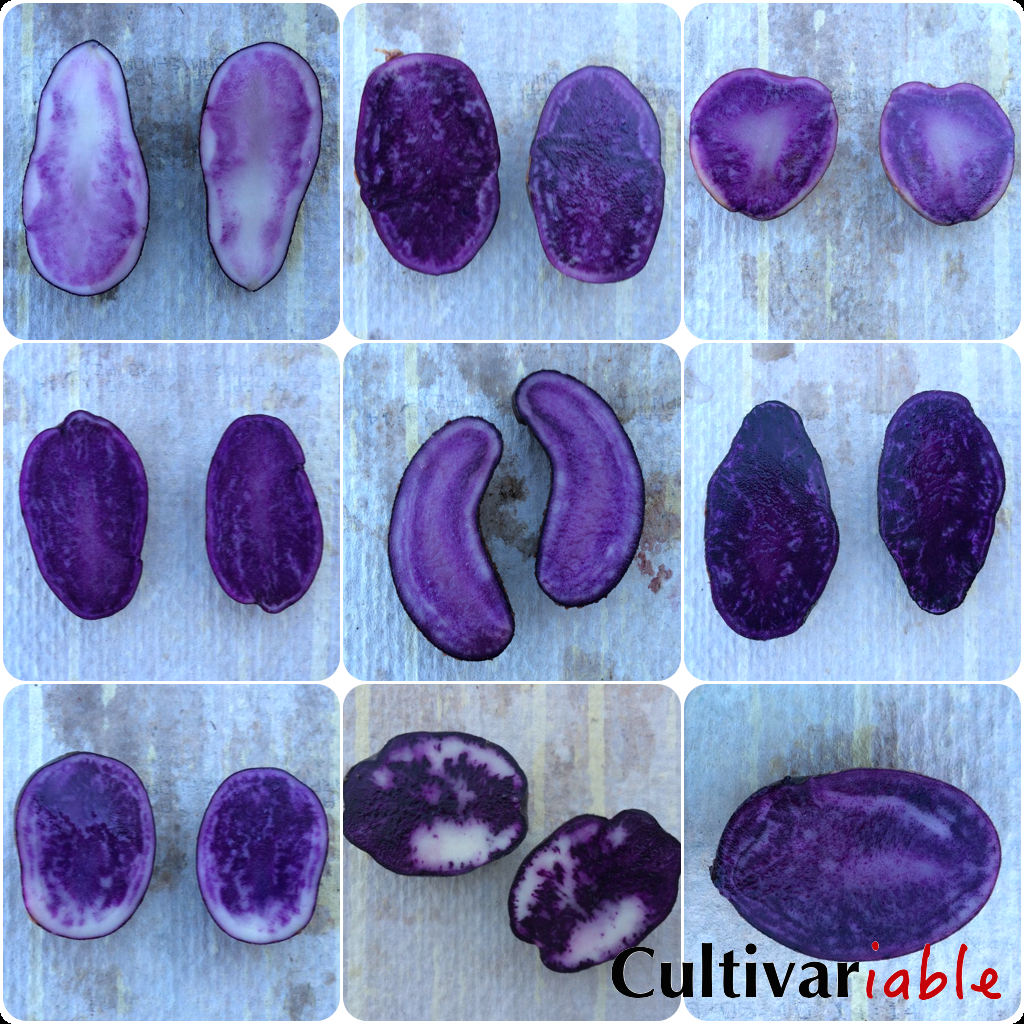

Purple and Red Pigments
The P locus controls production of petunidin, an anthocyanidin that produces purple coloration. The R locus controls production of pelargonidin, an anthocyanidin that produces red coloration. P is epistatic to R; when both are dominant, purple color is produced rather than red. This is thought to be due to competition between the two genes for a precursor involved in anthocyanidin production. (In general, potato produce only red or blue, but I have observed a few andigena type tetraploids that have both blue and red skin.) Like anthocyanins, anthocyanidin structure is affected by pH and temperature. At higher pH and/or temperature, color may not be as vibrant (or in some cases, not present at all). Purple pigments are found in some wild species, but only the domesticated potato has red pigments (Dodds 1962).
Purple and red pigments are distributed to the skin when I is dominant. In the case of homozygous recessive ii, purple and red pigments are not distributed to the skin even if P and/or R are dominant.
Purple and red pigments are distributed to the flesh when Pf is dominant. In the case of homozygous recessive pfpf, diploids do not distribute red or purple pigments to the tuber flesh. Little information is available about the operation of Pf in tetraploids.
A particularly dark purple pigment, malvidin, is found only in certain tetraploid potatoes, like Negresse (also known as Congo, but not to be confused with the Congo that is synonymous with All Blue). Simmonds (1965) hypothesized that this pigment was formed by a second methylation of petunidin and that this was controlled by a similar gene to Ac. He also noted that a petunia gene, K, performs the same function. Why this is only seen in tetraploids is uncertain. This could be another allele of Ac but is probably not a tetraploid dosage effect because it was produced in a dihaploid cross.
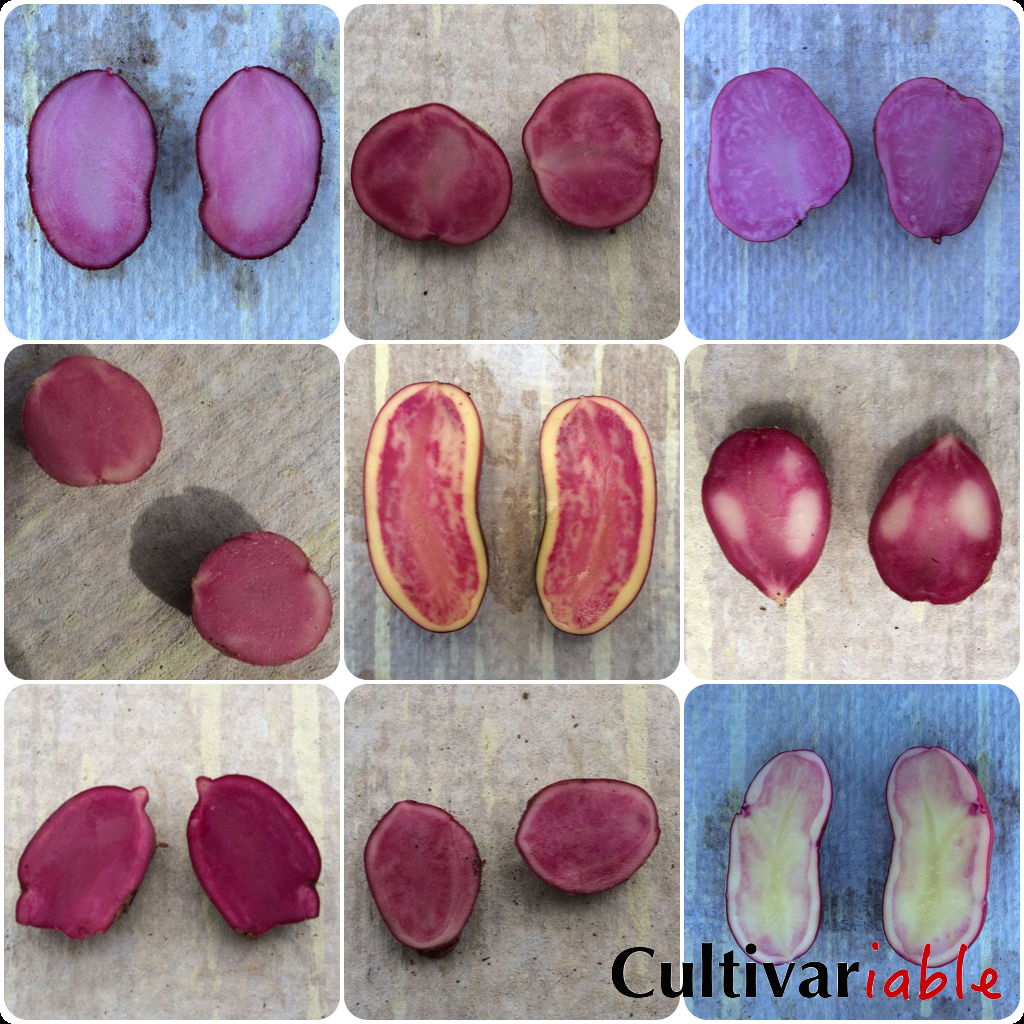

There are additional QTLs affecting red and purple flesh color. Loci that influence flesh color have been identified on chromosomes 5, 8, and 9. More detailed information about the operation of these loci has not yet been published as far as I am aware. Several recent papers have described a fairly large number of QTLs affecting tuber flesh color. Amateur breeders probably don’t need to be too concerned about this; the P-R-I complex gives reasonably good predictability. It is enough to be aware that there may be other genes at work if a cross does not conform well to expectations. With a better understanding of all of the involved genes, we could get better predictability of completeness and intensity of pigmentation, but it doesn’t seem like this information will be available in an easily digestible form any time soon.
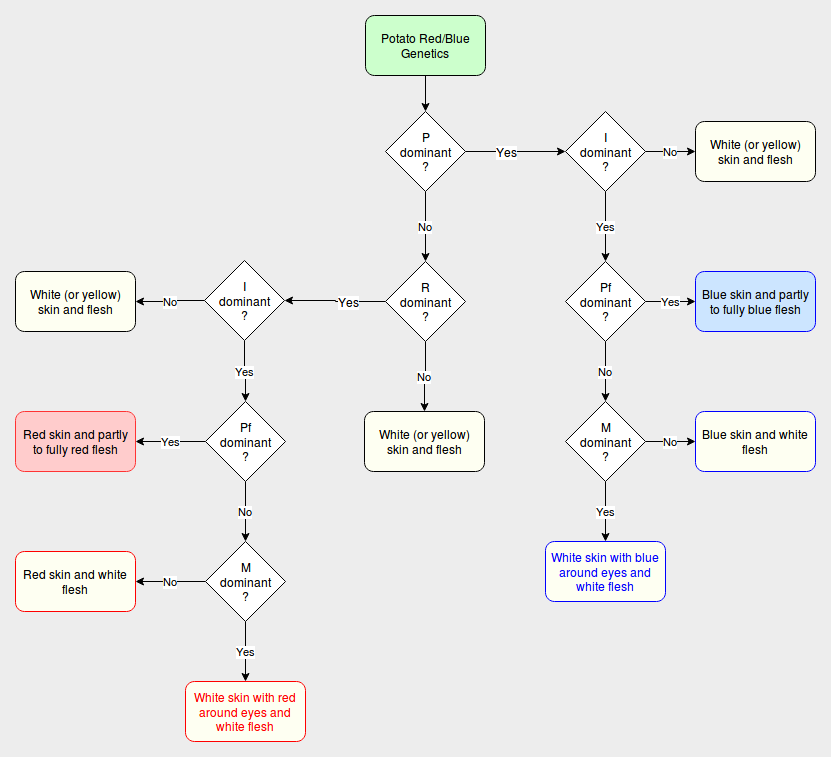
There is a flowchart showing red and blue phenotype outcomes at the bottom of this article.
Yellow and Orange Pigments
Yellow and orange flesh are fairly common in Andean potatoes, but not often seen in modern varieties. Because yellow skin and flesh often occur together, my assumption is that the same genes control yellow/orange skin and flesh, although I haven’t seen a single paper that addresses this. Historically, the multiallelic locus Y has been implicated in expression of both orange or yellow flesh. Or (orange flesh) is dominant over Y (yellow flesh), which is dominant over y (white flesh). Or favors production of zeaxanthin over lutein, Y lutein over zeaxanthin, and homozygous yy produces little of either, resulting in white flesh. Yellow flesh has also been found to be a quantitative trait with a number of different QTLs involved that affect intensity. I have searched, but not found any accounting of what occurs when both R or P and Or or Y are dominant. From experience, I know that yellow pigments and red or purple pigments can express together, but I don’t know the details. It is also not clear how the Y locus interacts with the various color distribution loci. Again, I know from experience that yellow or orange pigments can sometimes appear in the leaves, but I found no information indicating whether this is controlled by the distribution genes covered above or by different mechanisms. In practice, I have never seen a cross result that made it appear that orange flesh is dominant; instead, orange flesh almost always gives way to less intense yellow flesh.
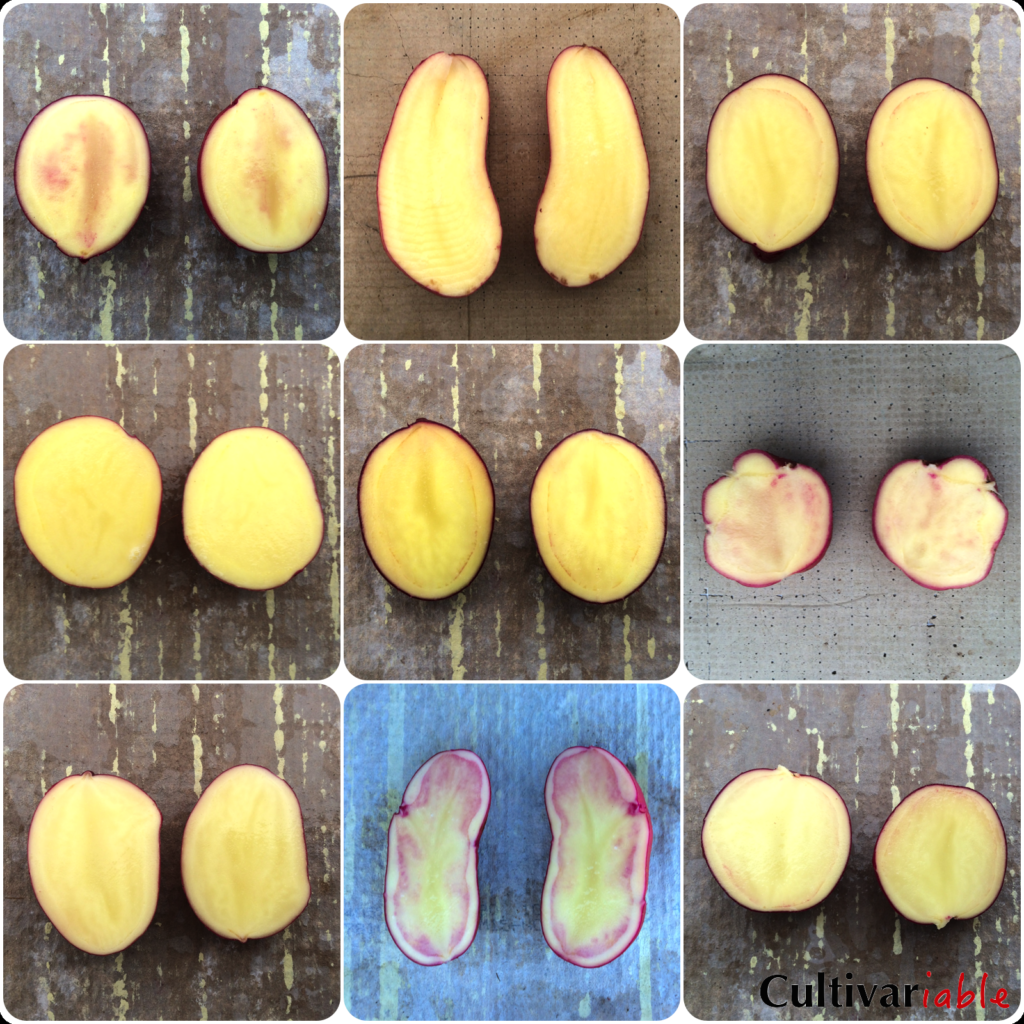

As noted above, including Or in this locus is now considered dubious. More recent information (Wolters 2010) indicates that orange is developed primarily through the combination of a homozygous recessive pairing of the Zep gene allele 1 (involved in zeaxanthin production) and Chy2 gene allele 3. Very few tetraploid potatoes contain Zep allele 1, which limits the possibility of orange flesh in tetraploids without bringing it from diploids.
Color Distribution
The B, I, and F complex controls distribution of pigments to many parts of the plant. These genes probably only distribute purple and red pigment, not orange and yellow, although I have not seen this stated explicitly. B, I, and F comprise a linkage group. They are often inherited together.
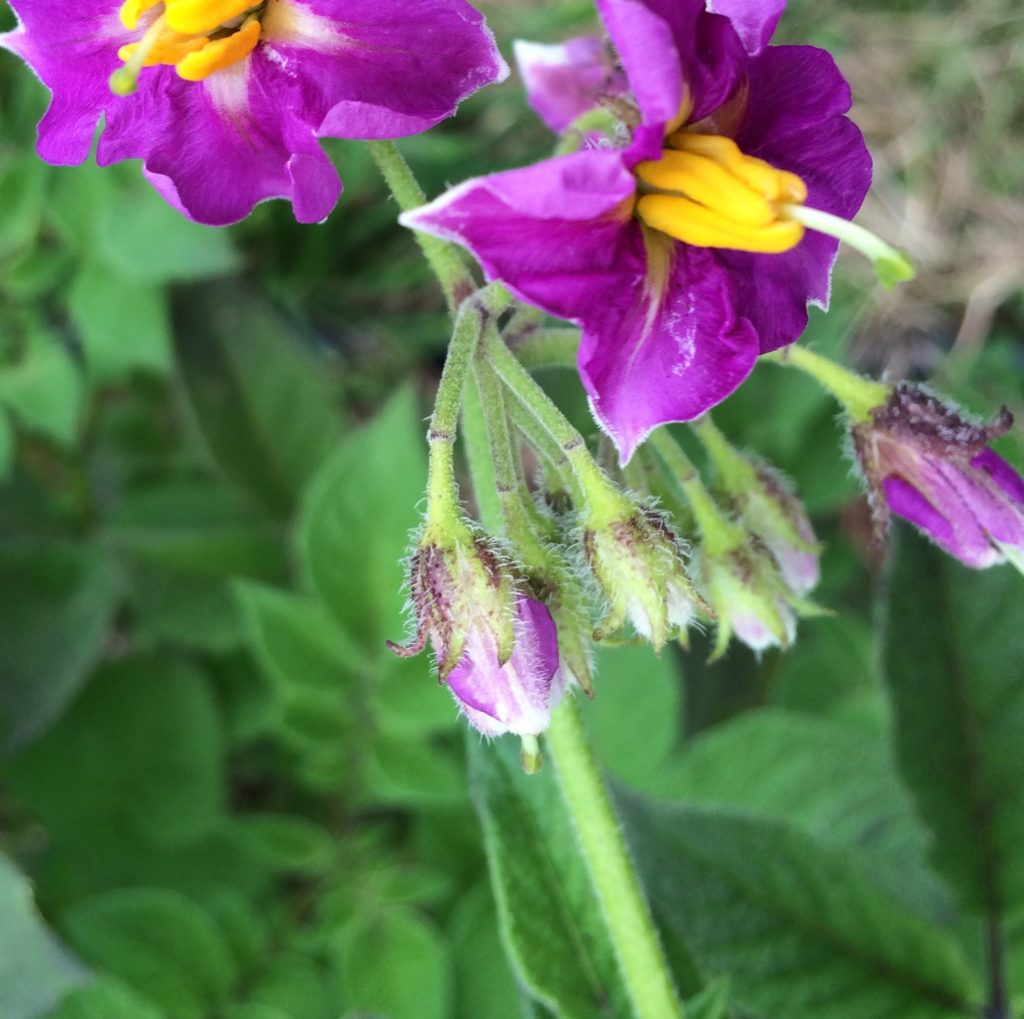
Ba-d distributes color to the flower abscission zone. This is the joint where the flower connects to the peduncle. There is usually just a small ring right at the joint. In the case of homozygous recessive bb, the joint is the same color as the rest of the inflorescence.
Bb-d distributes color to the tuber eyebrows – the arc shaped structure next to each eye. In the case of homozygous recessive bb, the eyebrow may not be colored when the rest of the skin is.
Bc-d distributes color to the seed spot or embryo spot. This is actually the cotyledonary node, where the hypocotyl is joined to the cotyledons. In the case of homozygous recessive bb, there is no seed spot. The seed spot is so named because you can see the pigmentation through the seed coat. This allows for screening for seeds that have this trait, which can be particularly useful when it is linked to other traits of interest.
Bd distributes color to the nodes on the stem and possibly also the tuber. In the case of homozygous recessive bb, the result appears to depend on whether the stem has red or blue color, in which case colorless bands can appear at the nodes. This may also extend into cleared spots around the eyes if the tuber skin is colored.
I (D) distributes color to the tuber skin and appears to be dosage dependent. For example, you would expect stronger color in an II diploid than in an Ii. Similarly, in tetraploids, the tubers of a plant with IIII would likely be much more strongly colored than a plant with Iiii. In diploids, I also controls expression of colored flesh (Pf). In the case of recessive ii, tubers will not have colored flesh even if Pf is dominant. Distribution of color to flesh in tetraploids is more complicated and I have not seen it clearly elucidated anywhere.
While these alleles serve as a reasonably good guide to the possibilities, the book is far from closed on this subject. For example, there are some Andean potatoes that are entirely white or yellow on the outside but have blue flesh. This combination does not fit with the options described above. It is good to keep in mind that most of the alleles described have not been determined through biochemical analysis; they are abstractions based on morphological observations and statistical analysis of progeny out of controlled crosses. In that sense, they may not be “real” even though they are useful.
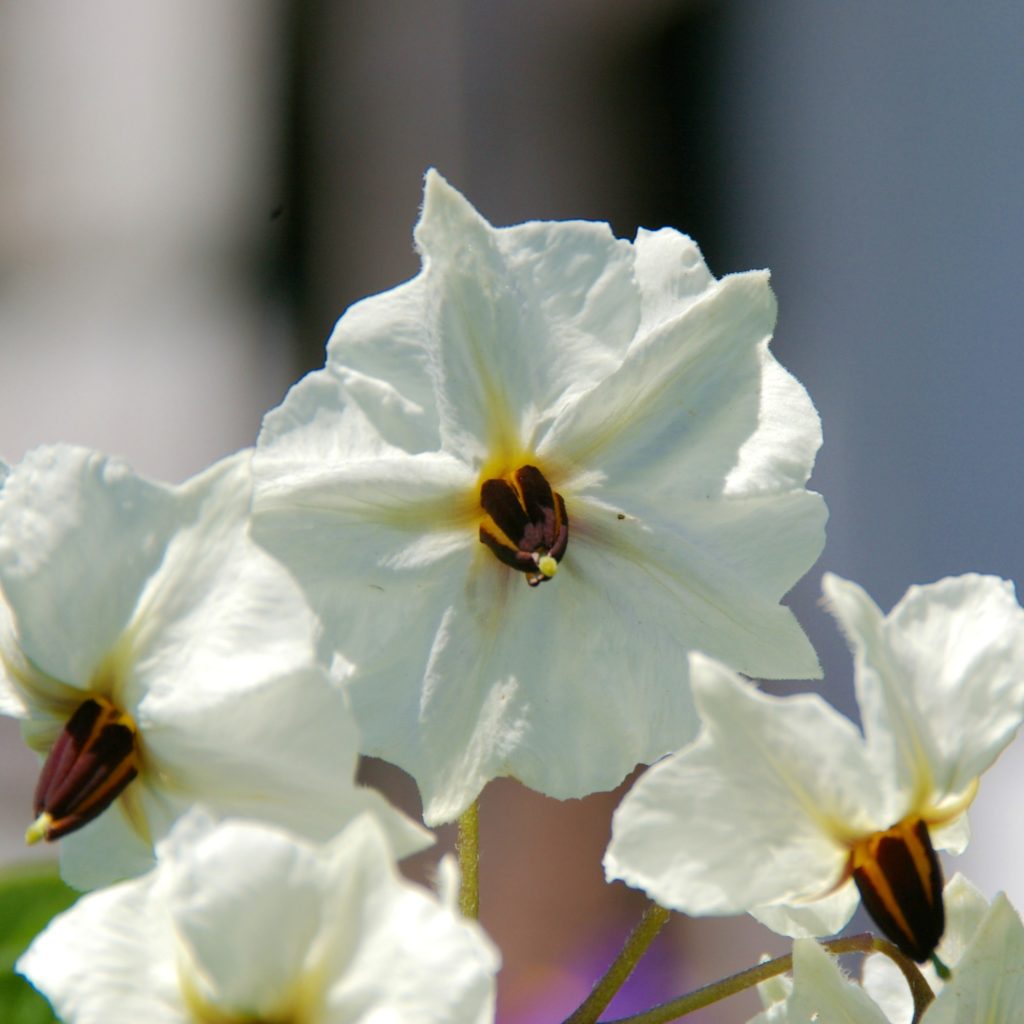
Flower Color
As noted above, F is a color distribution gene and it acts primarily on the distribution of red and purple pigments to the flowers, as is detailed in the following chart:
| Genotype | Phenotype | Notes | Examples | |
| R-ppF- | Pink corolla | Red/purple because only R is expressed. Additionally, the color of the flowers is reddish-purple, rather than pure red, because the gene R produces the cyanidin in the flowers vs pelargonidin in the tubers (Howard 1970). |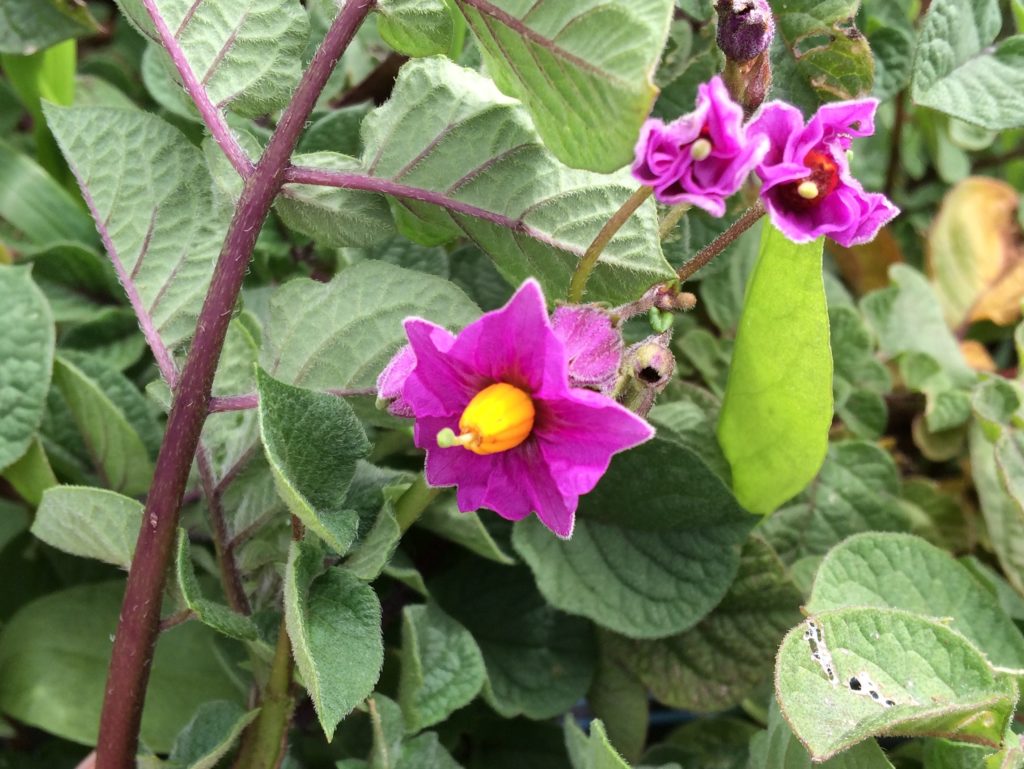  |
|
| rrP-F- | Blue corolla | Blue because only P is expressed. |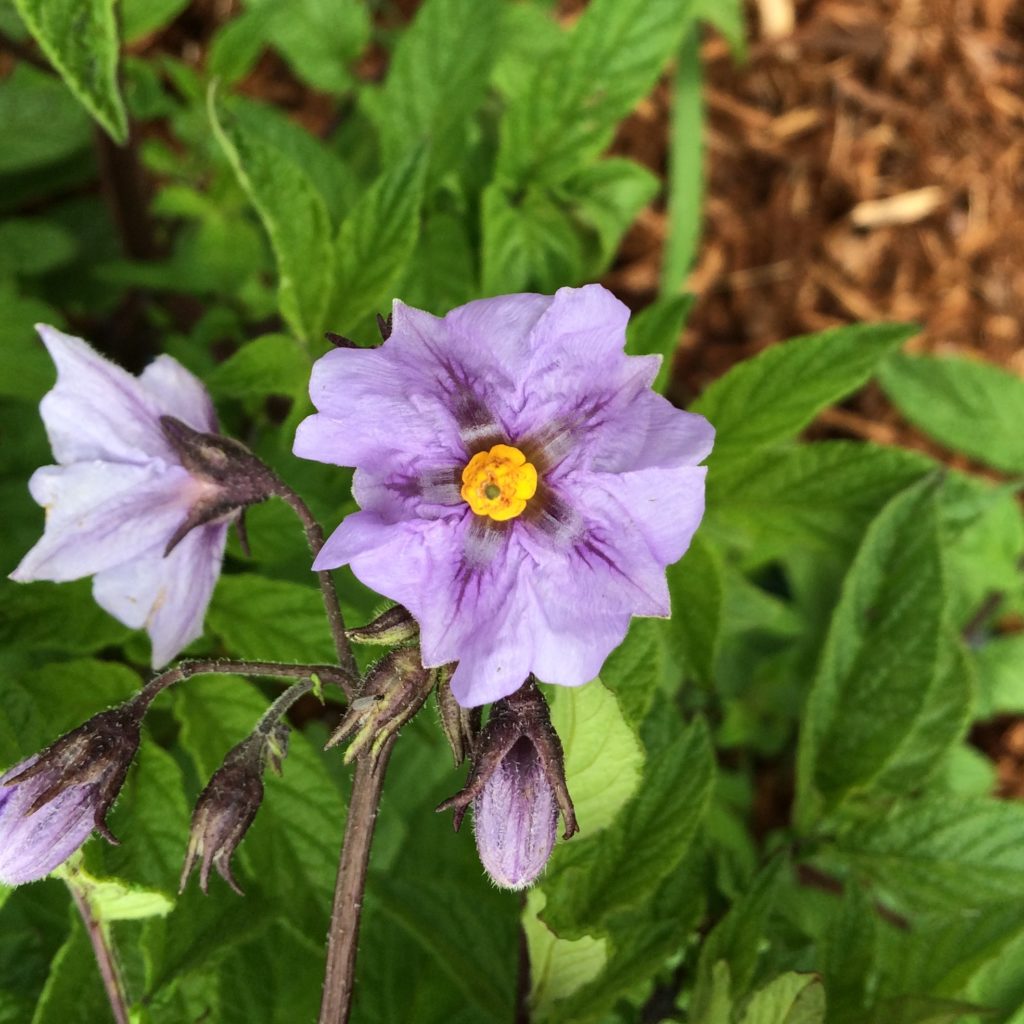  |
|
| R-P-F- | Purple corolla | Blue/purple because P is epistatic to R, but R is still able to contribute some red pigment. |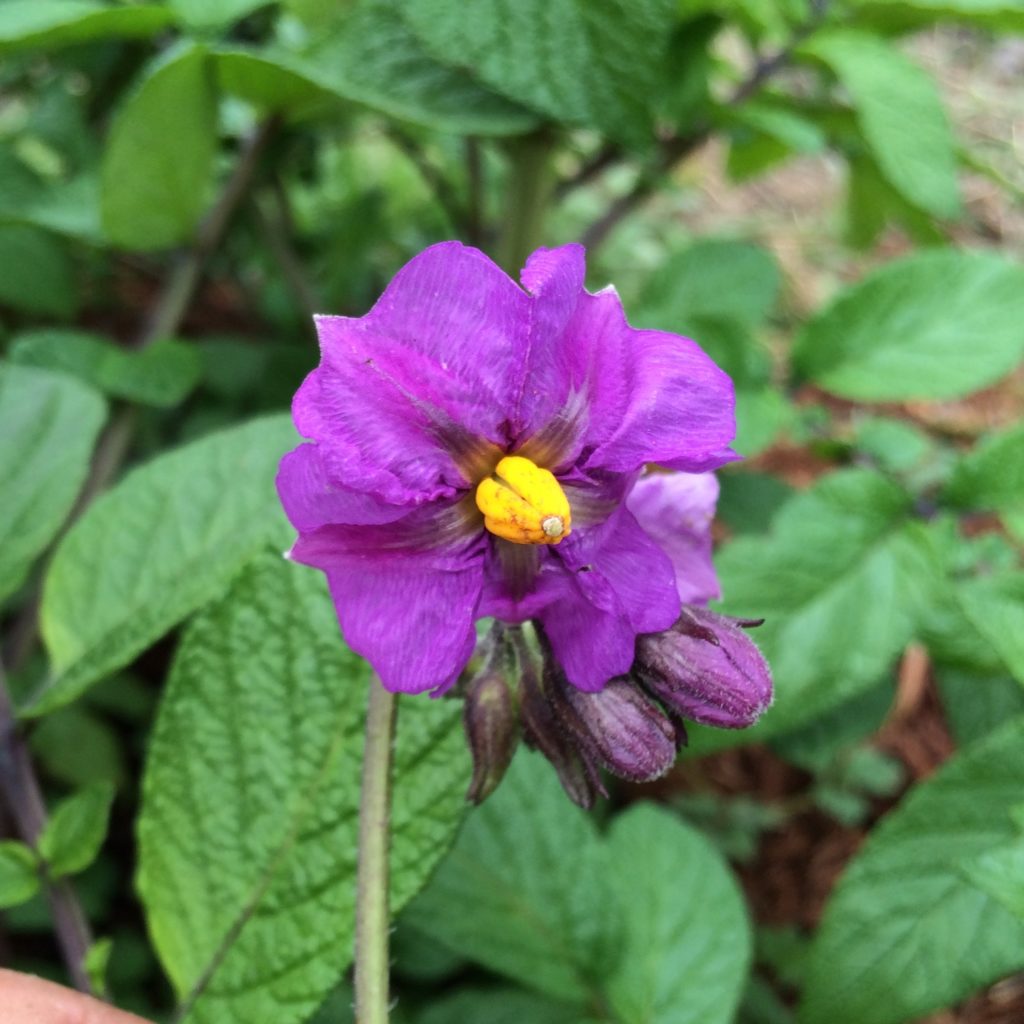  |
|
|
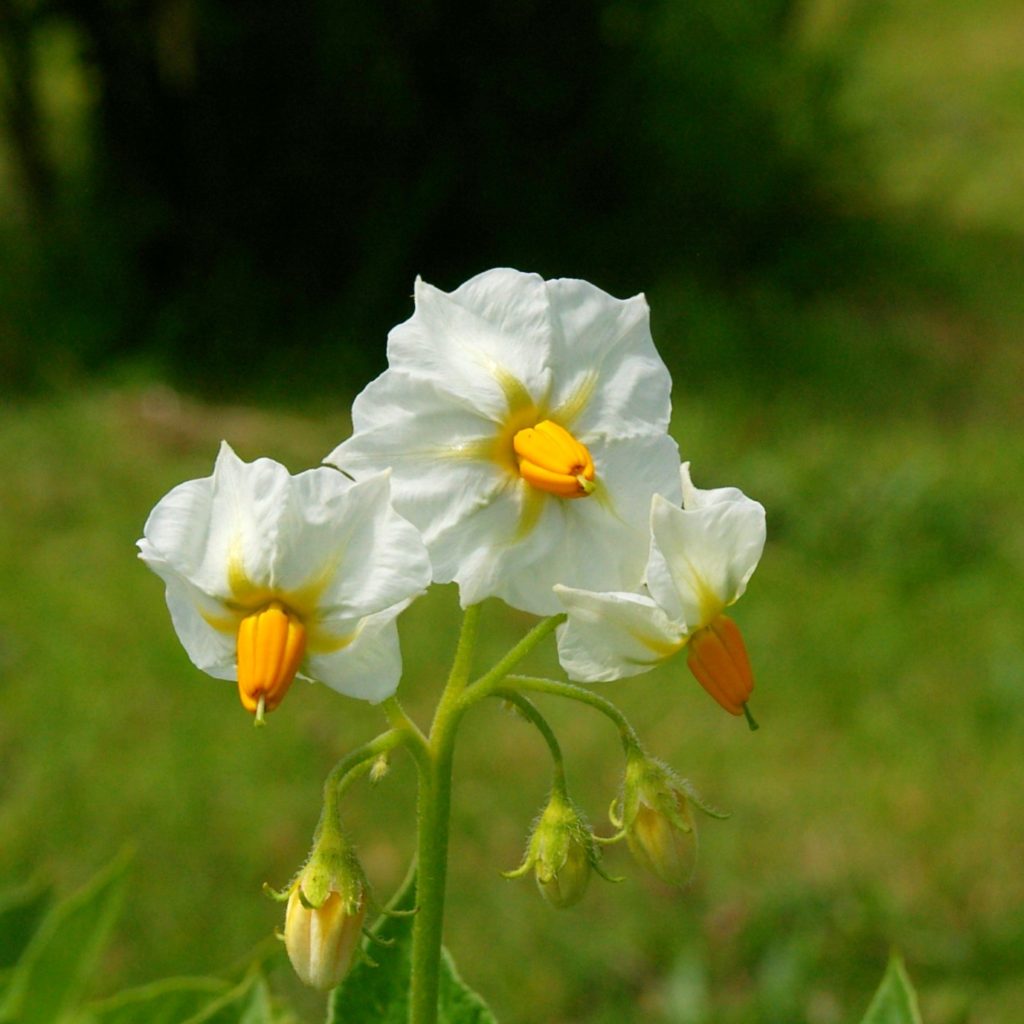
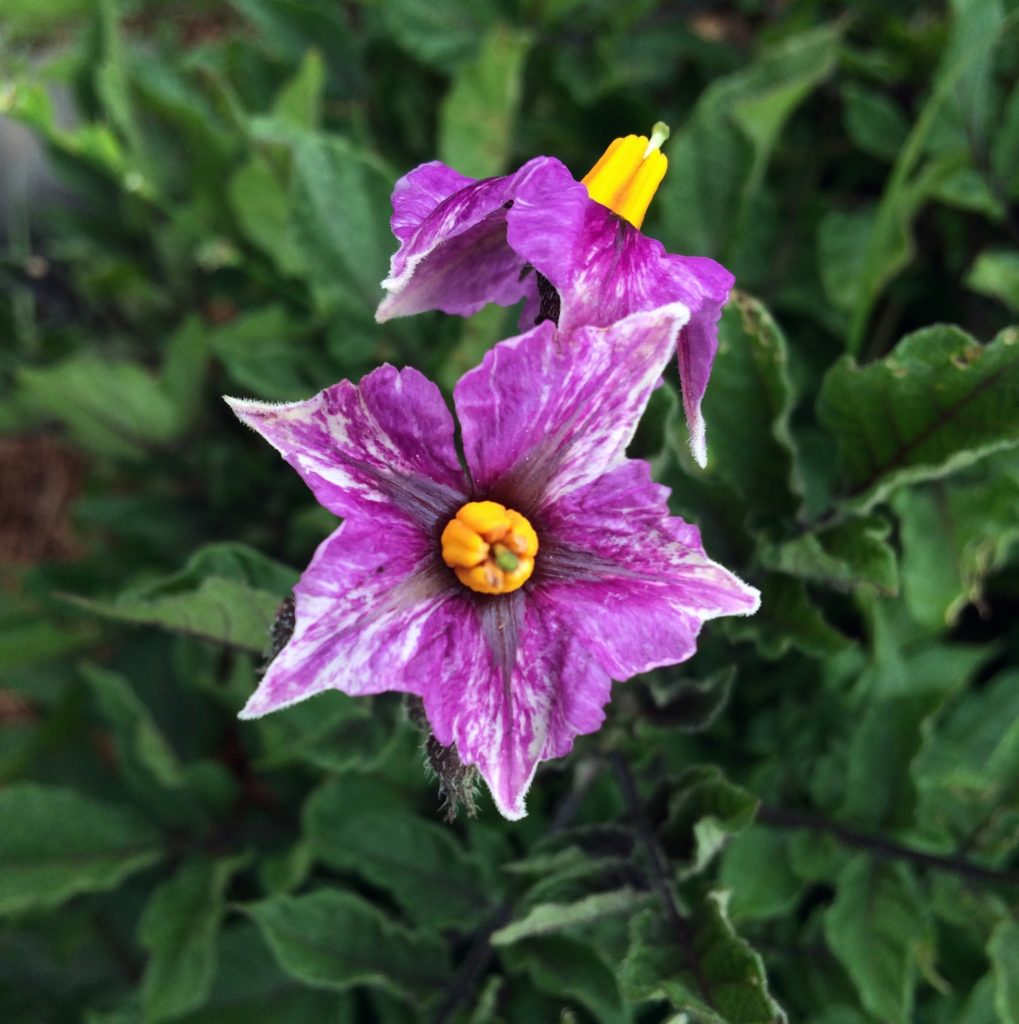
R-P-ff rrrrP-ff R-ppppff |
White corolla |
Usually white because ff blocks pigment distribution to the flower, but sometimes produces flecked flowers, perhaps due to interaction with another gene or a leaky allele. |
 |
 |
| rrppff | White corolla | White because ff blocks pigment distribution to the flower but also because there is no pigment produced. No possibility for flecked flowers, despite homozygous ff, because there is no pigment to distribute. |  |
|
| rrppF- | White corolla | White because there is no pigment to distribute to the flower. |  |
|
| RpwRpwP-F- | Pale blue corolla | The combination of P and homozygous Rpw produces pale blue flowers according to Dodds (1955). I’m not sure if I have seen this color. |
The chart applies to diploids, which are more straightforward to predict. Tetraploids have four copies of each allele, which can result in some intermediate phenotypes.
Very rarely, potato flowers can also be light yellow. I have found no information about the genetics of this flower color. It is appears to result from the partial conversion of petals to anthers. Some of the flowers on yellow petal cultivars occasionally convert fully to anthers, producing flowers with no petals. This is most commonly seen in Gelbblühender Stamm, an East German variety.
Although anther color is related to overall pigment production, I have not found any clear explanation of its inheritance. It is not included within the F locus. For example, I have several potatoes with red skin and flesh, but white corollas. That presumably puts them in the R-ppppffff genotype, but they also have red anthers, so ffff is clearly not blocking distribution of red to the anthers. Or, perhaps F is affected by dosage and these are something like Ffff. I think that potatoes with red anthers always have red flesh and potatoes with blue anthers always have blue flesh, but, unfortunately, the opposite is not always true. Potatoes with red or blue flesh and white corollas occasionally have yellow anthers. I have not yet seen this in combination with a pigmented abscission zone, so perhaps recessive b genotypes might explain the yellow anthers. That seems rather obvious, so it would probably have been discovered and published by now if it were true.
Foliage Color
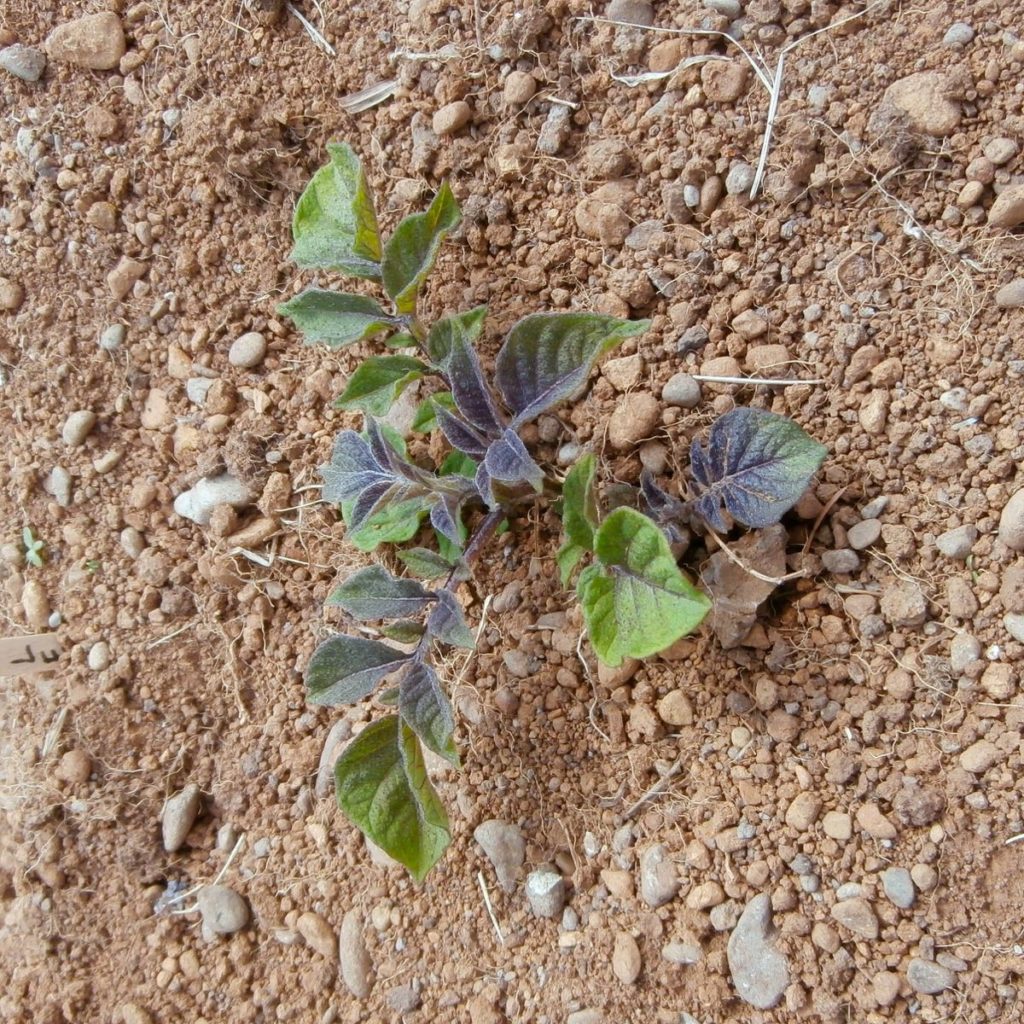
Most of the foliage color distribution genes, Pd, Pw, and Pv (Ul), have been described only for use with diploids. The same genes must function in tetraploids, but perhaps they do not express in a clear fashion or perhaps they have just not been sufficiently studied. I can say from experience that foliage color seems to be easier to predict in diploids than tetraploids.
Russeting
Russeting is more a condition of skin structure than color, but still probably fits better in this post than on the future post that I will do on tuber shape and structure. Unfortunately, the inheritance of russeting has not received much study. The only available study was done with diploids and suggests that russeting is controlled by three independently segregating genes. All three genes must be dominant for russeting to appear. The loci have not been named. Inheritance under a three gene system may be very difficult to sort out in tetraploids. Russeting may also appear as a defect in normally non-russet tubers in calcium poor soils.
Known Potato Color Genotypes
The following table lists some potatoes for which the color genotype has been fully or partially determined. These varieties may be useful for making test crosses to determine the color genetics of unknown varieties. I will try to add genotypes here whenever I run across them.
| Variety | Genotype | Phenotype |
| Prince Hairy | iiii rrrr P- | white skin/flesh, blue flowers |
| Chieftain | I- R- pppp | red skin, white flesh |
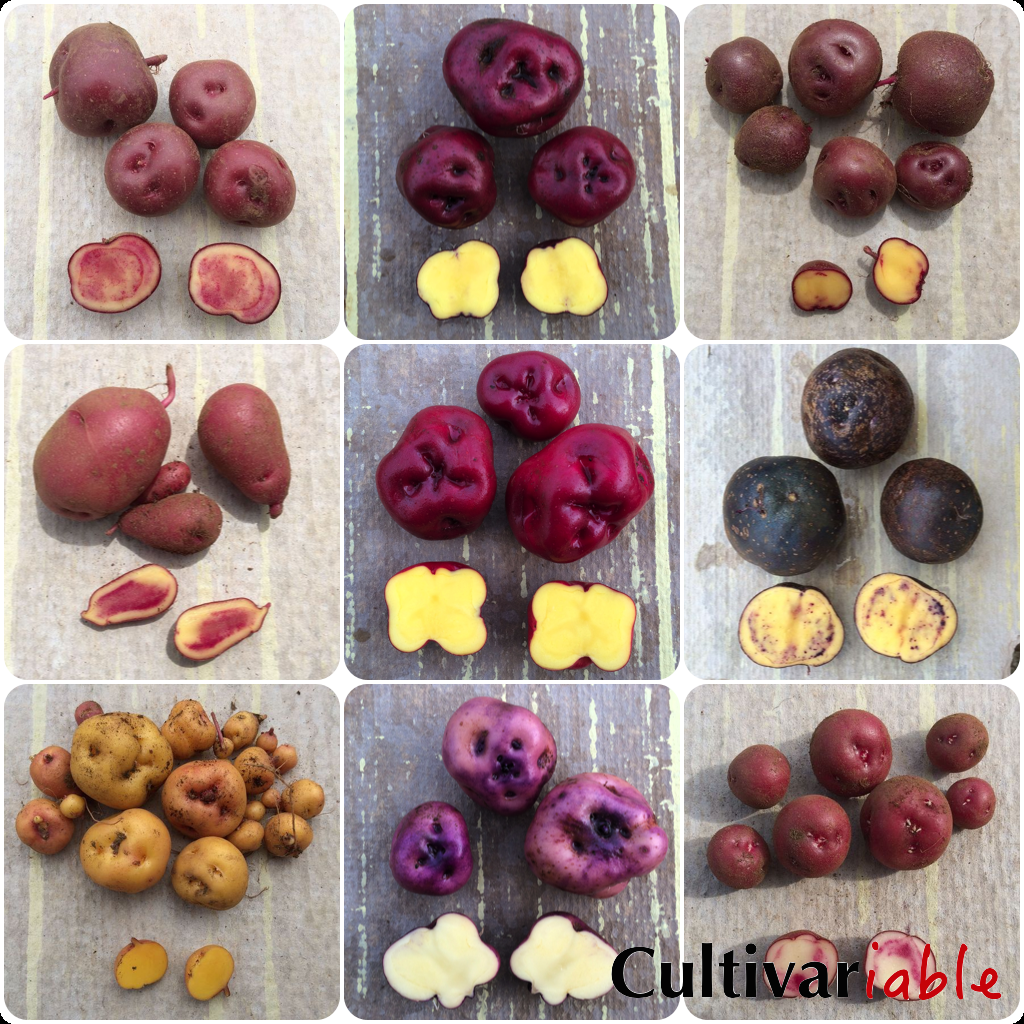
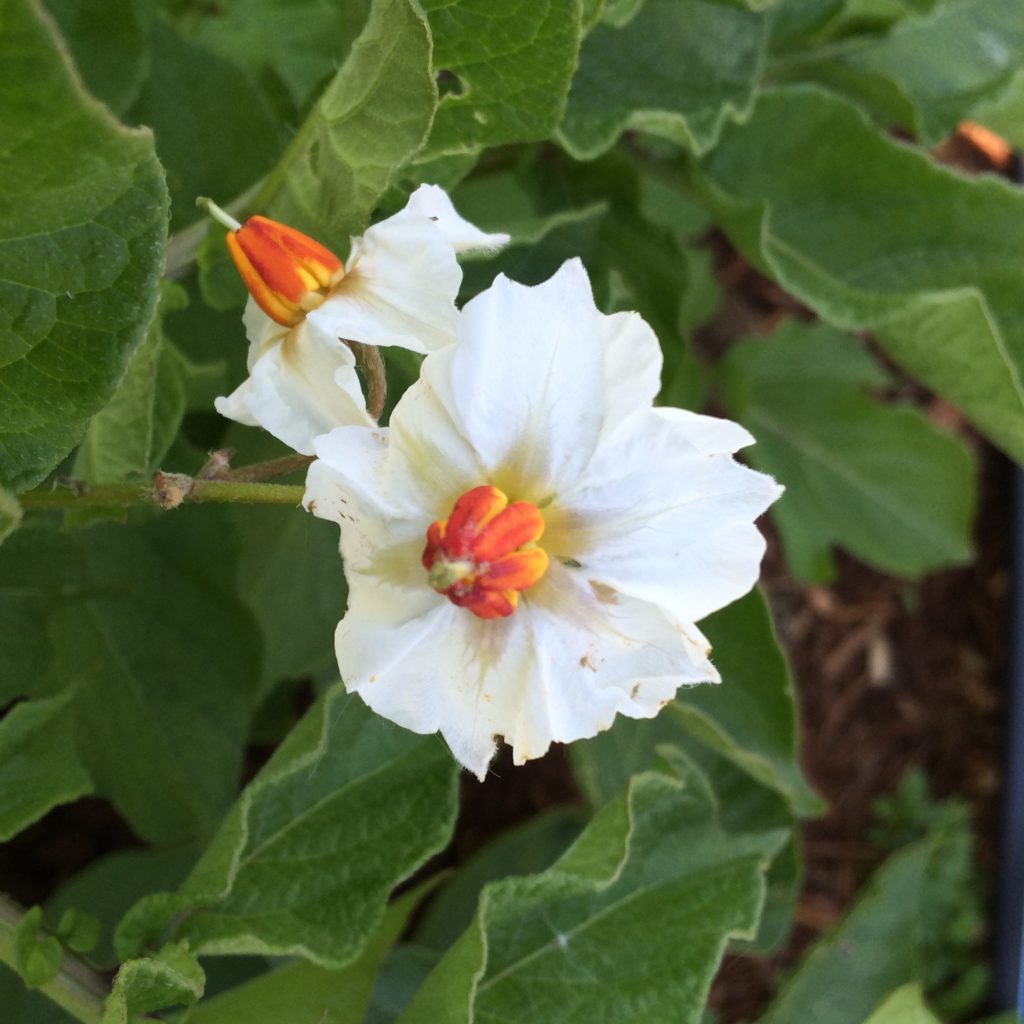
Potato Color Flowchart




Very informative. Thanks
I found this page because I started putting together a potato colour genetics chart like this. It was taking me some time because, as you said, this information is scattered across scientific literature. Looks like you beat me to it and did a great job.
I am so glad I found this page as this is an excellent resource (and there is no point reinventing the wheel). Thank you!
Is anybody have an idea why Red Norland would show white and red skin areas at harvest ?
The white is really white, and the red is also the good red color.
Could it be:
– impact of high temperature (soil and air) or drought on gene expression
– impact of drought (ie potatoes were not irrigated)
– impact of herbicides
– or a combination ?
I would need to see it, but the most likely explanation is a somatic mutation. Potatoes produce chimeras fairly frequently, where there is either a mutation that changes the color or exposure of a different layer of the epidermis that has different color. When these mutations are stable, they can be used to select a new variety, which is known as a “sport.” If there are any eyes in the white skin area, you should be able to start plants from those eyes that form only white tubers.
I have the same question and I am wondering if you ever got to the bottom of this?
Fascinating. “There’s no such thing as a red potato in the flesh, and why is that?” I said at the dinner table, and as usual I was wrong about what there is such a thing as. I see now there are some commercial red-fleshed varieties.
Nice and comprehensive summary of colour genetics! Perhaps you might clarify the difference between synthesis and regulation: Some genes are coding for enzymes involved in the biochemical synthesis of these pigments.
Some genes are coding for transcription factors that bind to the DNA in the nucleus and can switch on/off genes. Regulation of gene expression allow the plant to control in which tissues pigments are produced. Cells in roots carry the same DNA as in flowers, but pigmentation genes are never switched on.
Chy2 and Y are two names for the same thing on chromosome 10. Chy refers to the name of the enzyme (= beta-carotene hydroxylase), whereas Y refers to the name of the dominant phenotype (=yellow flesh).
Looking forward on your post on tuber shape. it is on chromosome 10 controled by OFP20 (https://www.nature.com/articles/s41467-018-07216-8)
Thanks very much for the suggestions! I didn’t make the Chy2/Y connection and that makes much sense now. The tuber shape article is languishing due to lack of information, unfortunately. I have substantially more phenotypes than genotypes to associate with them, so I am trying to sort that out.